hr_data |>
summarise(mean_hr = mean(hr))# A tibble: 1 × 1
mean_hr
<dbl>
1 85.3across()docs/cleaning.qmdIn the last session, we used regular expressions to add a id column to the HR-type data so that we have an identifier between the HR-type datasets and the survey data. By having this identifier, we’re one step closer to joining them together. But we still have several other things to process first. The next one we will do it process the HR-type data so that the date times (in seconds) match the time units in the survey data (in minutes). For that, we need to be able to summarise the HR-type data into a value per minute.
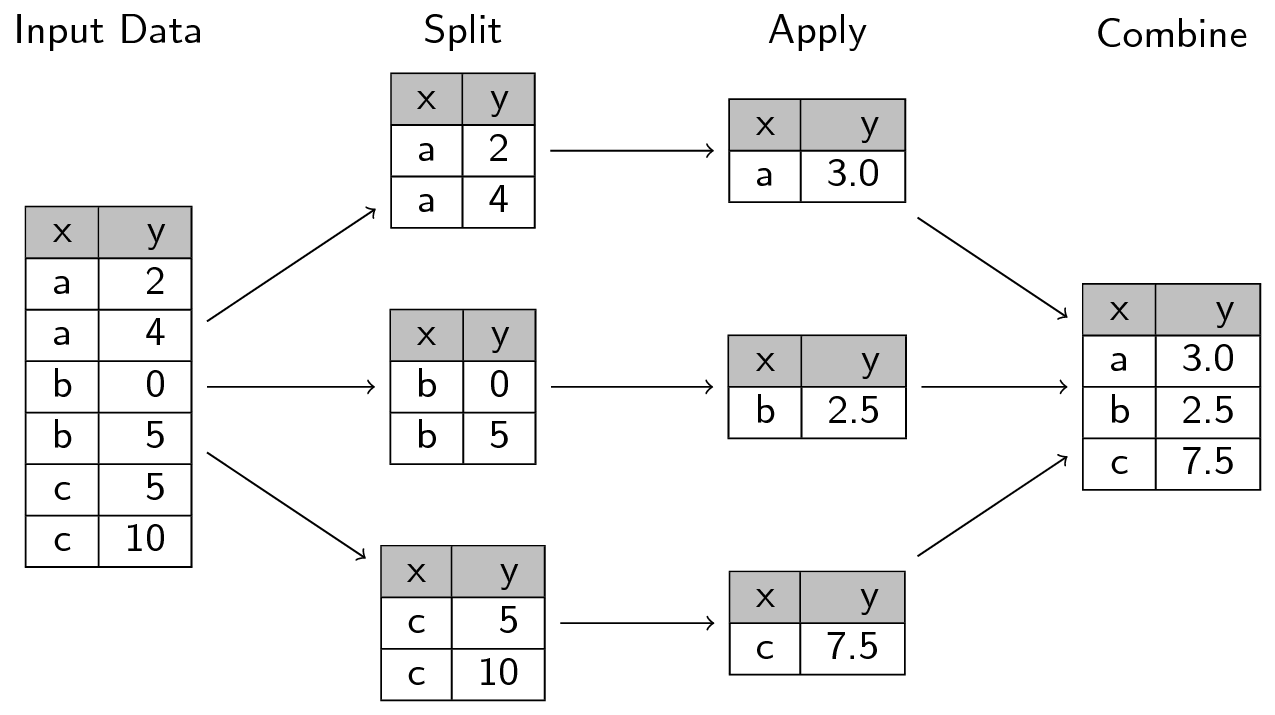
summarise(), its .by argument, and across() functions.Verbally cover this section before moving on to using across() while summarizing. Specifically, review the split-apply-combine technique via the image below as well as in the context of using summarise() on multiple columns and multiple functions.
Time: ~10 minutes.
We’re taking a quick detour to briefly talk about a concept that perfectly illustrates how vectorization and functionals fit into doing data analysis. The concept is called the split-apply-combine technique, which we covered in the beginner R workshop. The method is:
So when you split data into multiple groups, you create a list (or a vector) that you can then apply (e.g. with the map functional) a statistical technique to each group through vectorization, and where you finally combine (e.g. with join that we will cover later or with list_rbind()). This technique works really well for a range of tasks, including for our task of summarizing some of the data so we can merge it all into one dataset.
Functionals and vectorization are integral components of how R works and they appear throughout many of R’s functions and packages. They are particularly used throughout the tidyverse packages like dplyr. Let’s get into some more advanced features of dplyr functions that work as functionals.
There are many “verbs” in dplyr, like select(), rename(), mutate(), summarise(), and group_by() (covered in more detail in the introductory workshop). The common usage of these verbs is through acting on and directly using the column names (e.g. without " quotes around the column name). Like most tidyverse functions, dplyr verbs are designed with a strong functional programming approach. But many dplyr verbs can also be used as functionals like we covered in previous sessions with map(), where they take functions as input. For instance, summarise() uses several functional programming concepts: Create a new column using an action that may or may not be based on other columns and output a single value from that action. Using an example with our data, to calculate the mean of a column and create a new column from that mean for the glucose values, you would do:
hr_data |>
summarise(mean_hr = mean(hr))# A tibble: 1 × 1
mean_hr
<dbl>
1 85.3This is declarative because we ask R to give us the mean of hr using the mean() function and to output a data frame with only one row using summarise(). If we wanted to calculate another summary statistic, like the standard deviation, we would just add another line to the summarise() function:
# A tibble: 1 × 2
mean_hr sd_hr
<dbl> <dbl>
1 85.3 15.6Repeating this pattern works fine with a few columns and maybe one or two summary statistics. But what if you wanted to calculate the mean, standard deviation, the median, and maybe also the maximum and minimum values? And what if you wanted to this for several different columns? If you repeated the above code, that will be a lot of (annoying) repetition. That’s where the power of functional programming and applying functionals (also called higher-order functions) shows its strength.
In the functionals session in Chapter 18, you used map() to apply a function to a list of paths to import the data. In dplyr, there is a function that works almost exactly like map(), but is used specifically for applying functions to columns in a data frame. This function is called across().
Unlike in map(), where you have to give it a list that contains different objects, in across() you give it a vector of columns instead. Just like map(), you also give across() a function (as an object without ()) to apply to the columns.
While map() is a more general-purpose function, across() is specifically designed to only work in dplyr verbs like mutate() or summarise() and within the context of a data frame.
When you’re ready to continue, place the sticky/paper hat on your computer to indicate this to the teacher 👒 🎩
across()
Open up ?across in the Console and go over the arguments with the learners. Go over the first two arguments again, reinforcing what they read.
Also, before coding again, remind everyone that we still only import the first 100 rows of each data file. So if some of the data itself seems weird, that is the reason why. Remind them that we do this to more quickly prototype and test code out.
Before we start using across(), let’s look at the help page for it. In the Console, type ?across and hit enter. This will open the help page for across(). For this workshop, we will only go over the first two arguments. The first argument is the columns you want to work on. You can use c() to combine multiple columns together. The second argument is the function you want to apply to those columns. You can use list() to combine multiple functions together.
Let’s try out using across() on the hr_data. Just to confirm that we’re using the same data, let’s read in the HR data and run get_participant_id() on it again. So, go to the bottom of your docs/learning.qmd file and create a new header called ## Summarising with across and create a code chunk below that with Ctrl-Alt-ICtrl-Alt-I or with the Palette (Ctrl-Shift-PCtrl-Shift-P, then type “new chunk”).
docs/learning.qmd
hr_data <- read_all("HR.csv.gz") |>
get_participant_id()Then, below that, we will do a very simple example of using across() to calculate the mean of the hr values.
docs/learning.qmd
hr_data |>
summarise(across(hr, mean))# A tibble: 1 × 1
hr
<dbl>
1 85.3This is nice, but let’s try also calculating the median. The help documentation of across() says that you have to wrap multiple functions into a list(). Let’s try that:
docs/learning.qmd
hr_data |>
summarise(across(hr, list(mean, median)))# A tibble: 1 × 2
hr_1 hr_2
<dbl> <dbl>
1 85.3 82.2It works, but, the column names are not helpful. The column names are hr_1 and hr_2, which doesn’t tell us which is which. We can add the names of the functions by using a named list. In our case, a named list would look like:
They don’t need to write this code in the Console, just explain to them what the code means and does, while showing the output. Same with the code below with the anonymous function.
Console
list(mean = mean)$mean
function (x, ...)
UseMethod("mean")
<bytecode: 0x5652865f9840>
<environment: namespace:base># or
list(average = mean)$average
function (x, ...)
UseMethod("mean")
<bytecode: 0x5652865f9840>
<environment: namespace:base># or
list(ave = mean)$ave
function (x, ...)
UseMethod("mean")
<bytecode: 0x5652865f9840>
<environment: namespace:base>See how the left hand side of the = is the name and the right hand side is the function? This named list is what across() can use to add the name to the end of the column name. Just like with map() we can also give it anonymous functions if our function is a bit more complex. For instance, if we needed to remove NA values from the calculations, we would do:
For now, we don’t need to use anonymous functions. Let’s all try using a named list with only one function, like list(mean = mean):
docs/learning.qmd
hr_data |>
summarise(across(hr, list(mean = mean)))# A tibble: 1 × 1
hr_mean
<dbl>
1 85.3Then, we can add more functions to the named list. Let’s add the median and standard deviation:
docs/learning.qmd
hr_data |>
summarise(
across(
hr,
list(mean = mean, sd = sd, median = median)
)
)# A tibble: 1 × 3
hr_mean hr_sd hr_median
<dbl> <dbl> <dbl>
1 85.3 15.6 82.2The great thing about using across() is that you can also use it with all the tidyselect functions in the first argument. In our case, we have datasets with columns that have different names. So if we made this code into a function, it would only work for datasets that have a hr column. That’s not very useful. Instead, we can use tidyselect functions to tell across() to apply the functions to columns based on some condition. Since all of our HR-type data have columns that are numeric, we can use the where() function with is.numeric (notice the lack of ()). Let’s try it out by replacing hr with where(is.numeric):
docs/learning.qmd
hr_data |>
summarise(
across(
where(is.numeric),
list(mean = mean, sd = sd, median = median)
)
)# A tibble: 1 × 3
hr_mean hr_sd hr_median
<dbl> <dbl> <dbl>
1 85.3 15.6 82.2Running that code gives us the same output as before, but now it’s generic enough that we can use it with any other the other datasets. Neat 😁
But, having only one row for the summary statistic for us isn’t very useful. Summarising is very powerful when combined with grouping. Which is what we’d like to do in order to effectively join the data. But before we can do that, we need to talk about dates and times.
Working with datetime data can be surprisingly difficult and tricky. There’s timezones, daylight savings time, leap years, and different date formats that all can cause a dozen different problems. On top of that, but datetimes can have nanosecond and second level precision. In R, there is a wonderful package called lubridate that helps with working with dates and times. It can’t solve all date and time problems, but it can help with a many of them. And, thankfully, lubridate is part of the tidyverse ecosystem, so we don’t need to explicitly load it.
In the docs/learning.qmd file, at the bottom of the document create a new code chunk called ## Working with dates and below that create a new code chunk with Ctrl-Alt-ICtrl-Alt-I or with the Palette (Ctrl-Shift-PCtrl-Shift-P, then type “new chunk”). We’ll use mutate() again to create a column called datetime, as well as .before to put it before the other columns.
Our collection_datetime column is already in a datetime format, but it is in seconds, while the survey data is in minutes. So, we need to convert the seconds into minutes. There are many ways to do this, but the easiest is to use the round_date() function from lubridate. This function rounds a datetime to the nearest unit of time. So, we can use it to round the collection_datetime to the nearest minute. The round_date() function takes two arguments: the datetime column and the unit of time to round to (e.g. “minute”, “hour”, “day”, etc.). It can even be used to round to the nearest 5 minutes (by using "5 mins") or 30 seconds (with "30 secs"). Let’s round to the nearest minute by using round_date() with the unit argument set to "minute":
hr_data |>
mutate(
collection_datetime = round_date(collection_datetime, unit = "minute")
)# A tibble: 57,183 × 3
id collection_datetime hr
<chr> <dttm> <dbl>
1 15 2020-07-07 16:43:00 83.7
2 15 2020-07-07 16:43:00 78.2
3 15 2020-07-07 16:44:00 72.5
4 15 2020-07-07 16:44:00 73.2
5 15 2020-07-07 16:44:00 77.8
6 15 2020-07-07 16:45:00 83.7
7 15 2020-07-07 16:45:00 84.1
8 15 2020-07-07 16:45:00 85.2
9 15 2020-07-07 16:45:00 83.7
10 15 2020-07-07 16:45:00 83.1
# ℹ 57,173 more rowsOur next step is to summarise by this collection_datetime column, so we have the measurement values per minute rather than per second. First, let’s add lubridate as a package dependency:
Console
usethis::use_package("lubridate")We used summarise() with the functional across() to calculate summary statistics for columns based on a condition for the “apply-combine” part. But now, we can do the “split” part for the datetimes by minute. In dplyr, one way to use the split-apply-combine technique is by using the summarise() function with its .by argument.
You can also pipe group_by() into summarise(), but you have to remember to use ungroup() afterwards. Using group_by() can be useful if you want to do multiple actions on the grouped dataset, since dplyr functions mutate() and summarise() also work on grouped data. At the end, you’d always have to ungroup() and if you forget, it can give wrong output that you might not realise.
If you only want to do a single action on groups, you can instead use the .by argument in summarise() or in mutate(). This is a more concise way of doing the group_by() and ungroup() workflow, since the grouping only happens in the summarise() or mutate() function. You can learn more about .by in the help page by running ?dplyr_by in the Console.
Let’s combine what we did with round_date() and what we did using summarise() and across(). At the bottom of the docs/learning.qmd file, create a new header called ## Summarising by groups and create a code chunk below that with Ctrl-Alt-ICtrl-Alt-I or with the Palette (Ctrl-Shift-PCtrl-Shift-P, then type “new chunk”). Rather than create a new column for a variable called datetime, let’s replace the existing collection_datetime column with the rounded version. And at the end of the summarise() function, let’s add the .by argument and set it to both id and collection_datetime within c(). This will split the data up by the id and collection_datetime columns, apply the summary statistics to each group, and then combine the results back together to then output that new data frame.
docs/learning.qmd
# A tibble: 12,785 × 5
id collection_datetime hr_mean hr_sd hr_median
<chr> <dttm> <dbl> <dbl> <dbl>
1 15 2020-07-07 16:43:00 81.0 3.83 81.0
2 15 2020-07-07 16:44:00 74.5 2.89 73.2
3 15 2020-07-07 16:45:00 83.8 0.744 83.7
4 15 2020-07-07 16:46:00 81.7 3.11 81.7
5 15 2020-07-07 16:47:00 76.7 2.50 75.6
6 15 2020-07-07 16:48:00 83.0 NA 83.0
7 15 2020-07-07 16:49:00 85.8 0.575 85.9
8 15 2020-07-07 16:50:00 88.0 2.48 87.8
9 15 2020-07-07 16:51:00 91.4 NA 91.4
10 15 2020-07-07 16:52:00 90.7 1.69 90.4
# ℹ 12,775 more rowsThe .by argument can be used similar to how select() is used, so you can use tidyselect functions to select the columns you want to group by. If you wanted to group by more than one column, you’d wrap the columns in c(). For our purposes, we don’t need to do that.
Amazing! Before continuing to the exercise, let’s run styler with the Palette (Ctrl-Shift-PCtrl-Shift-P, then type “style file”) on both docs/learning.qmd and on R/functions.R. Then we will render the Quarto document with Ctrl-Shift-KCtrl-Shift-K or with the Palette (Ctrl-Shift-PCtrl-Shift-P, then type “render”) to confirm that everything runs as it should. If the rendering works, switch to the Git interface and add and commit the changes so far with Ctrl-Alt-MCtrl-Alt-M or with the Palette (Ctrl-Shift-PCtrl-Shift-P, then type “commit”) and push to GitHub.
Time: ~15 minutes.
It’s your turn to convert the code into a function so that you can use it on the other datasets. It should be able to work like this for the HR data:
hr_data <- read_all("HR.csv.gz") |>
get_participant_id() |>
summarise_by_datetime()You should be able to do the same thing for the other datasets (IBI, TEMP, BVP, and EDA) using this function.
Here’s a scaffold of the function to help you get started:
function(), add one argument called data.hr_data with the data argument.summarise_by_datetime() function to its package with ::. There should be three packages used within the function.usethis::use_package().docs/learning.qmd with the Palette (Ctrl-Shift-PCtrl-Shift-P, then type “style file”).summarise_by_datetime(hr_data). It should give you the same output as before.R/functions.R.Add the package dependency for tidyselect:
usethis::use_package("tidyselect")The function should look like this in the R/functions.R file:
R/functions.R
#' Summarise values in a data frame by a rounded datetime
#'
#' @param data A data frame with at least a `collection_datetime` column and
#' some numeric columns to summarise.
#'
#' @returns A summarised data frame.
#'
summarise_by_datetime <- function(data) {
summarised_data <- data |>
dplyr::mutate(
collection_datetime = lubridate::round_date(
collection_datetime,
unit = "minute"
)
) |>
dplyr::summarise(
dplyr::across(
tidyselect::where(is.numeric),
list(mean = mean, sd = sd, median = median)
),
.by = c(id, collection_datetime)
)
return(summarised_data)
}And you should be able to use it like this in either the Console or in the docs/learning.qmd file:
hr_data <- read_all("HR.csv.gz") |>
get_participant_id() |>
summarise_by_datetime()When you’re ready to continue, place the sticky/paper hat on your computer to indicate this to the teacher 👒 🎩
docs/cleaning.qmd
Time: ~15 minutes.
You’re now at a point where you have functions to read in and process all of the HR-type data. So let’s start filling in the docs/cleaning.qmd file. This is the main document you’ll use to keep all the code that will clean and process the data into a single dataset. So, open up the docs/cleaning.qmd file (you can open files quickly with Ctrl-.Ctrl-.).
setup code chunk, add the source(here::here("R/functions.R")) line to load in the functions you have created so far.docs/cleaning.qmd, create a new header called ## Prepare HR data. Create a new code chunk below that with Ctrl-Alt-ICtrl-Alt-I or with the Palette (Ctrl-Shift-PCtrl-Shift-P, then type “new chunk”).read_all(), get_participant_id(), and summarise_by_datetime() to read in and process the HR data by piping them together with |>. Assign the output to a new variable called hr_data.ibi_data, temp_data, bvp_data, and eda_data respectively.In later exercises, you’ll come back to this docs/cleaning.qmd file to add more code to continue processing the data and eventually join it all together.
Just remember that we are only importing the first 100 rows of each dataset, so if it is confusing why data may look a bit different or that it’s too small, this is the reason why. We do this to more quickly prototype and test code out.
docs/cleaning.qmd
hr_data <- read_all("HR.csv.gz") |>
get_participant_id() |>
summarise_by_datetime()
ibi_data <- read_all("IBI.csv.gz") |>
get_participant_id() |>
summarise_by_datetime()
temp_data <- read_all("TEMP.csv.gz") |>
get_participant_id() |>
summarise_by_datetime()
bvp_data <- read_all("BVP.csv.gz") |>
get_participant_id() |>
summarise_by_datetime()
eda_data <- read_all("EDA.csv.gz") |>
get_participant_id() |>
summarise_by_datetime()When you’re ready to continue, place the sticky/paper hat on your computer to indicate this to the teacher 👒 🎩
Time: ~10 minutes.
If you’ve finished the previous exercises early and want to do something a bit harder, try out this exercise. You might have noticed that the code you wrote above to read in each of the HR-type datasets is very similar. The only thing that changes is the filename. So, you can create a function that takes in a filename as an argument and then reads in the data, adds the participant id, and summarises it by datetime. This function should be able to work with any of the HR-type datasets. Name the new function read_sensor_data().
If you did the extra exercise in one of the previous sessions and added the max_rows argument to the read_all() function, add that argument to this new function as well and set it to 100 by default. Make sure to pass this argument to the read_all() function within the body of the new function.
Make sure to follow workflows we’ve been using throughout this workshop: styling the R code, testing the code, committing, and pushing to GitHub. Because this is an optional, extra exercise, we don’t include a solution for this exercise.
R/functions.R
read_sensor_data <- function(filename) {
data <- read_all(filename) |>
get_participant_id() |>
summarise_by_datetime()
return(data)
}
# Or if you added the max_rows argument to read_all():
read_sensor_data <- function(filename, max_rows = 100) {
data <- read_all(filename, max_rows = max_rows) |>
get_participant_id() |>
summarise_by_datetime()
return(data)
}When you’re ready to continue, place the sticky/paper hat on your computer to indicate this to the teacher 👒 🎩
Time: ~8 minutes.
If you’ve finished the previous exercises early and want to do something a bit harder, try out this exercise. Modify the summarise_by_datetime() function in the R/functions.R file so that you can set your own unit of time to round to. For instance, you might want to round to the nearest 5 minutes or 30 seconds. You can do this by adding a new argument to the function called unit and then using that argument in the round_date() function. This new argument should have a default value of "minute" so that when we eventually join the datasets together later on, than you don’t have issues with the datasets not matching up.
Make sure to add this new argument to the read_sensor_data() function as well, so you can set the unit of time from there as you read in the data.
Make sure to follow the workflows we’ve been using throughout this workshop: styling the R code, testing the code, committing, and pushing to GitHub. Because this is an optional, extra exercise, we don’t include a solution for this exercise.
R/functions.R
read_sensor <- function(filename, max_rows = 100, unit = "minute") {
data <- read_all(filename, max_rows = max_rows) |>
get_participant_id() |>
summarise_by_datetime(unit = unit)
return(data)
}When you’re ready to continue, place the sticky/paper hat on your computer to indicate this to the teacher 👒 🎩
Time: ~5 minutes.
If you’ve finished the previous exercises and the optional extra exercise and there’s some time left, try out this exercise if you want to try something harder. Modify the summarise_by_datetime() in the R/functions.R file so that you can set your own summarising functions. To make it challenging, we won’t give you more hints aside from showing you how this function might look like when you use it:
Make sure to follow workflows we’ve been using throughout this workshop: styling the R code, testing the code, committing, and pushing to GitHub. Because this is an optional, extra exercise, we don’t include a solution for this exercise.
Good luck! 🎉
When you’re ready to continue, place the sticky/paper hat on your computer to indicate this to the teacher 👒 🎩
Quickly cover this and get them to do the survey before moving on to the discussion activity.
summarise(), its .by argument, and across() to use the split-apply-combine technique when needing to do an action on groups within the data.round_date().Please complete the survey for this session:
This lists some, but not all, of the code used in the section. Some code is incorporated into Markdown content, so is harder to automatically list here in a code chunk. The code below also includes the code from the exercises.
hr_data <- read_all("HR.csv.gz") |>
get_participant_id()
hr_data |>
summarise(across(hr, mean))
hr_data |>
summarise(across(hr, list(mean, median)))
list(mean = mean)
# or
list(average = mean)
# or
list(ave = mean)
list(mean = \(x) mean(x, na.rm = TRUE))
hr_data |>
summarise(across(hr, list(mean = mean)))
hr_data |>
summarise(
across(
hr,
list(mean = mean, sd = sd, median = median)
)
)
hr_data |>
summarise(
across(
where(is.numeric),
list(mean = mean, sd = sd, median = median)
)
)
hr_data |>
mutate(
collection_datetime = round_date(collection_datetime, unit = "minute")
)
usethis::use_package("lubridate")
hr_data |>
mutate(
collection_datetime = round_date(
collection_datetime,
unit = "minute"
)
) |>
summarise(
across(
where(is.numeric),
list(mean = mean, sd = sd, median = median)
),
.by = c(id, collection_datetime)
)
usethis::use_package("tidyselect")
#' Summarise values in a data frame by a rounded datetime
#'
#' @param data A data frame with at least a `collection_datetime` column and
#' some numeric columns to summarise.
#'
#' @returns A summarised data frame.
#'
summarise_by_datetime <- function(data) {
summarised_data <- data |>
dplyr::mutate(
collection_datetime = lubridate::round_date(
collection_datetime,
unit = "minute"
)
) |>
dplyr::summarise(
dplyr::across(
tidyselect::where(is.numeric),
list(mean = mean, sd = sd, median = median)
),
.by = c(id, collection_datetime)
)
return(summarised_data)
}
hr_data <- read_all("HR.csv.gz") |>
get_participant_id() |>
summarise_by_datetime()
hr_data <- read_all("HR.csv.gz") |>
get_participant_id() |>
summarise_by_datetime()
ibi_data <- read_all("IBI.csv.gz") |>
get_participant_id() |>
summarise_by_datetime()
temp_data <- read_all("TEMP.csv.gz") |>
get_participant_id() |>
summarise_by_datetime()
bvp_data <- read_all("BVP.csv.gz") |>
get_participant_id() |>
summarise_by_datetime()
eda_data <- read_all("EDA.csv.gz") |>
get_participant_id() |>
summarise_by_datetime()